This study presents an approach, for identifying and categorizing diseases in mango leaves expanding its scope beyond diseases while differentiating between healthy leaves and the impacted by these ailment. Using a process the proposed method involves preparing leaf images by adjusting their size converting them to grayscale and reducing noise with a filter. Subsequently K means segmentation is employed to areas affected by disease. This segmentation step is followed by processing using wavelet decomposition to enhance features in the images. An important aspect is the extraction of features where a Histogram of Oriented Gradients (HOG) is utilized to capture characteristics of the segmented regions. These features act as a representation for classification. The classification stage utilizes a Support Vector Machine (SVM) with a kernel showcasing its efficiency in distinguishing leaves from those affected by Anthracnose. Experimental findings demonstrate the accuracy of the proposed approach achieving precision, recall rates and F1 Score, for disease categories. The effectiveness of this methodology is illustrated through an assessment using a dataset containing images of healthy mango leaves as those affected by disease. The new method doesn't just help with spotting diseases but sets the stage for future progress, in precision farming and crop care.
Mango cultivation, globally recognized for its role in biodiversity and as a major source of fruits, has garnered increasing attention for promoting sustainable agricultural practices. However, the thriving mango plantations face a formidable challenge in the form of various diseases, including Mango malformation, Anthracnose, Bacterial flower disease, Golmachi, Moricha disease, and others. These diseases significantly impact mango trees worldwide, leading to substantial economic losses and hindering fruit production.
Plant diseases pose a severe threat to countries heavily reliant on agriculture. Pakistan, a significant contributor to global crop production, faces economic repercussions due to declining fruit quality and yields caused by these diseases. Identifying and addressing diseased plants through computer vision and image processing [1] techniques has become imperative. Deep learning techniques, particularly convolutional neural networks (CNNs) [2], have shown remarkable success in plant disease classification, offering a promising avenue for disease detection.
The mango, a staple fruit in Pakistan's agricultural industry, is susceptible to diseases like powdery mildew, sooty mold, anthracnose, and apical necrosis. Recognizing the economic impact of these diseases, researchers are exploring computer-based solutions for early disease detection. Traditional methods, relying on naked-eye observation, are costly and time-consuming. The complexity of mango plant structures and patterns necessitates efficient and robust techniques for automatic and accurate disease identification.
In this context, the proposed research leverages computer vision, image processing, and machine learning [3] methodologies to detect mango plant diseases at their initial stages. The research utilizes digital and mobile cameras to capture baseline images for analysis. By advancing the automated identification of mango diseases, this work aims to provide farmers with a tool to safeguard crops, reduce economic losses, and contribute to the sustainable growth of the agricultural sector.
In contemporary agricultural practices, the effective identification and mitigation of plant diseases stand as critical factors for sustaining global food production. Machine Learning, as an integral component of Artificial, emerges as a key player in this domain, facilitating automated systems to deliver precise and reliable results [4]. Inspired by real-world objects, Machine Learning techniques are designed to decipher complex patterns and variations inherent in the diverse landscape of plant diseases [5].
A multifaceted strategy is employed in the research approach. Wavelet transforms are used for improved feature extraction after K-means segmentation [6] of the mango leaf images. The most significant element of this approach is feature extraction, namely with the use of the Histogram of Oriented Gradients (HOG) [7]. A Support Vector Machine (SVM) classifier, an essential part of the pipeline for disease diagnosis, is trained using these features as its framework.
Mia et al.'s (2020) [8] study uses a Neural Network Ensemble (NNE) and Support Vector Machine (SVM) to tackle the problem of disease detection in mango plants. The suggested method, which focused on four distinct mango leaf diseases Dag, Golmachi, Moricha, and Shutimold disease achieved an average accuracy of 80%. To improve illness diagnosis, the authors used a variety of data manipulations, including zoom levels and rotations. The study highlights how machine learning may be used to automate disease detection, save farmers' time, and boost mango production all of which will help meet demand in the worldwide market. The literature study emphasizes how important their strategy is for improving farming methods and spurring economic expansion.
The limitations of manual detection rendered automated mango leaf disease recognition imperative, according to a study by Saleem et al. (2021) [9]. They obtain an astonishing 95.5% accuracy with a cubic support vector machine with their innovative "leaf vein-seg" segmentation, which integrates vein patterns and canonical correlation analysis-based fusion. The research employs a self-collected dataset to examine powdery mildew and sooty mold, and it proposes recommendations for future developments, highlighting the need for larger datasets and the use of innovative deep-learning [10] algorithms for improved disease detection.
Rehman et al. (2022) [11] present a novel deep transfer learning method for categorizing citrus diseases. The suggested approach makes use of two pre-trained models with transfer learning and feature fusion, MobileNetv2 and DenseNet201, using image augmentation and hybrid contrast stretching. The Whale Optimization Algorithm is used to optimize the feature set. This improves disease identification in citrus plants by exceeding current techniques and producing a 95.7% classification accuracy for six citrus diseases.
A Multilayer Convolutional Neural Network (MCNN) is presented by Singh et al. (2019) [12] to classify mango leaves infected with anthracnose. The work shows how well MCNN can identify between healthy and diseased leaves, having been validated on a dataset of 1070 images. The proposed approach outperforms other innovative techniques with a remarkable 97.13% classification accuracy. This novel method, which combines deep learning and computer vision, can identify anthracnose in mango trees early and effectively, which may reduce economic losses and improve crop quality.
Sujatha et al. (2021) assess the effectiveness of deep learning (DL) approaches (Inception-v3, VGG-16, VGG-19) with machine learning (ML) methods (Support Vector Machine, Random Forest, Stochastic Gradient Descent) for the diagnosis of citrus plant diseases [13]. According to the experimental data, DL approaches perform better than ML methods; VGG-16 achieves the greatest accuracy of 89.5% in disease classification. The research highlights the potential integration of artificial intelligence, specifically deep learning (DL), with the Internet of Things, cloud computing, and big data to improve agricultural decision-making.
To overcome the challenges associated with real-time crop monitoring, Usharani et al. [14] suggest a ground bot that comes with a camera and SVM algorithm to accurately detect pests. Their work is notable for automating the process of detecting crop and soil conditions using color-coded maps. Farmers are notified via phones and receive correct results on an LCD screen as a result of the integration of SVM processing at the server backend. With its insights into pest presence and ability to increase productivity with little computational time, this work is essential for real-time agricultural field monitoring.
Pradipkumar and Alagu Raja [15] offer a method utilizing machine learning (ML) from UAV images to address the need for effective tree species identification. They use the Histogram of Oriented Gradient (HOG) descriptor for feature extraction and classifiers including SVM, KNN, DT, RF, and XGBoost. HOG is evaluated in the study to other descriptors, such as Linear Binary Pattern (LBP) and Gray-Level Co-occurrence Matrix (GLCM). By integrating the estimations of many classifiers, XGBoost achieves the best accuracy (97.94%). This method demonstrates how machine learning and image processing may be used to automatically identify different tree species, which can help preserve biodiversity and be applied to a range of social issues.
Recent advances in computer vision further emphasize the importance of robust feature representations under challenging conditions. STRACK (2025) [16] introduced a framework for tracking small objects in low-light environments by focusing on invariant feature extraction and adaptive processing. Although STRACK targets object tracking, its emphasis on handling illumination variability and low-contrast regions is highly relevant to plant disease imagery, where lighting conditions often vary significantly.
Similarly, the HAMOT framework (2025) [17] proposed a hierarchical adaptive architecture for robust multi-object tracking in complex environments. HAMOT demonstrates that structured pre-processing and hierarchical feature modeling substantially improve robustness under visually challenging conditions. These findings indicate a growing trend toward multi-scale analysis and adaptive feature representation in visual recognition tasks.
Recently, Kamal et al. (2025) [18] proposed a hybrid framework combining the Meyer Wavelet Transform and a Jaccard Deep Q-Network (JDQN) for small object classification using multi-modal images. The study demonstrated that wavelet-based frequency decomposition significantly enhances discriminative feature representation, particularly for subtle and low-contrast objects. By integrating wavelet features with reinforcement learning, the proposed method achieved robust classification performance under complex imaging conditions. Although the approach is designed for multi-modal and small object classification tasks, it highlights the effectiveness of wavelet-based multi-scale analysis, which directly motivates the use of wavelet transforms for texture enhancement in plant disease classification.
While existing studies predominantly rely on deep learning architectures or single-stage machine learning models, many of them suffer from high computational cost, limited interpretability, and strong dependence on large-scale labeled datasets, which restrict their applicability in real-world agricultural environments. Moreover, limited attention has been given to integrated hybrid frameworks that systematically combine segmentation, multi-scale texture enhancement, and handcrafted feature extraction specifically for mango leaf disease classification under varying imaging conditions.
Motivated by these limitations, the present work proposes a computationally efficient and interpretable hybrid framework that integrates K-means segmentation, wavelet-based multi-scale enhancement, HOG feature extraction, and SVM classification. By jointly leveraging spatial segmentation and frequency-domain texture refinement, the proposed approach achieves high classification accuracy with reduced computational complexity, making it well suited for practical and resource-constrained agricultural applications.
Highlights
- Integrated Approach for Mango Leaf Disease Detection.
- Wavelet Transform & HOG Features to improve disease classification.
- SVM Classifier achieves 99.50% accuracy for mango leaf classification.
In this research, a comprehensive approach for mango leaf disease detection using a combination of image processing and machine learning techniques is presented. The dataset used in this proposed method was obtained from Kaggle website and consists of 2000 images, with 500 images per class of four categories: Healthy, Powdery Mildew, Sooty Mould and Anthracnose [19]. The research was carried out using MATLAB version 2020a. To ensure uniformity, the collected images with varying sizes are resized to the size of the HOG feature size which is 64 × 64 pixels. Some examples of datasets are shown in the Figure 1.
The initial stage of our proposed technique focuses on optimizing the input images for subsequent analysis.
3.1. Pre-Processing
This section presents the proposed mango leaf disease classification framework. Figure 2 illustrates the overall workflow of the proposed method, which consists of image acquisition, preprocessing, segmentation, wavelet-based refinement, feature extraction, and classification. The framework is designed to progressively enhance disease-relevant information while suppressing background noise, enabling accurate and efficient classification.
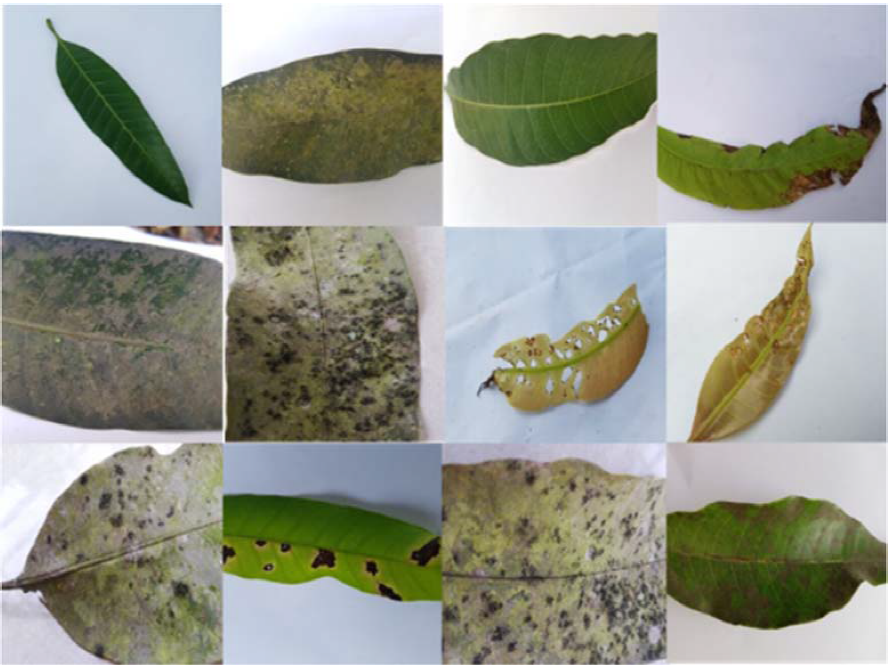

Figure 1. Example for Mango leaf dataset.
To establish uniformity in the dataset, collected RGB images are subjected to resizing operations. Each image is resized to a standardized dimension of 64 × 64 pixels. This step ensures consistent input sizes, laying the foundation for effective processing and feature extraction. The RGB images, now of uniform dimensions, undergo grayscale conversion [20]. This transformation is crucial for simplifying subsequent computations. The conversion, results in a single-channel grayscale representation of the original image.
To enhance the quality of the images and reduce noise interference, a median filter is applied. The grayscale image is processed, effectively replacing each pixel value with the median value of its local neighbourhood. This denoising step prepares the images for more robust segmentation and feature extraction. The pre-processing phase establishes a standardized and noise-reduced foundation for the subsequent stages of our proposed method. These initial steps are pivotal in ensuring the input images are appropriately formatted for effective disease detection in mango leaves. These pre-processing steps ensure standardized input conditions for subsequent segmentation. The workflow of the proposed work is depicted in Figure 2.
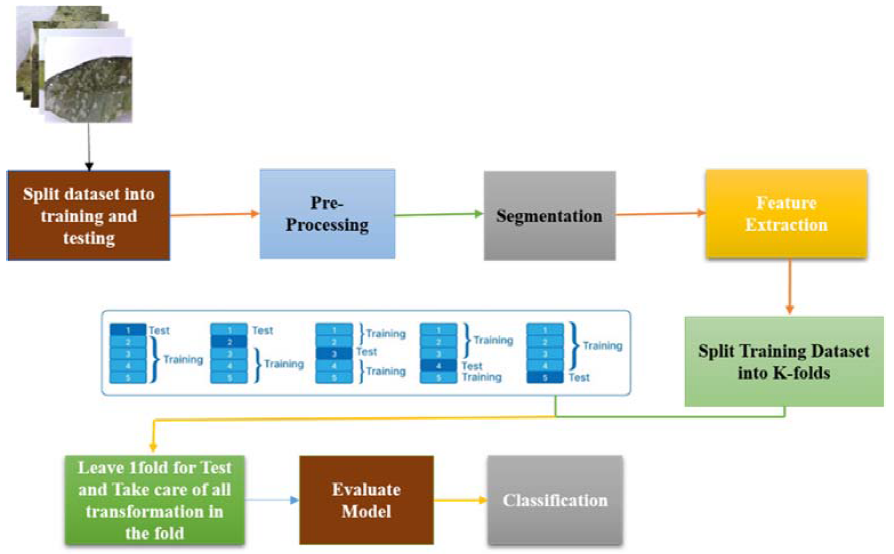
Figure 2. Workflow of the proposed work.
3.2. K-Means Based Leaf Segmentation
In the proposed methodology, a robust segmentation technique is employed to accurately delineate and isolate regions of interest within mango leaf images. Segmentation is a pivotal step in the image processing pipeline, facilitating the extraction of meaningful features for subsequent analysis. The algorithm encompasses a multi-step process, integrating K-means clustering and post-processing methods to enhance the precision of the segmentation results.
These pre-processing steps ensure standardized input conditions for subsequent segmentation. The resized and denoised images are then subjected to K-means clustering, a powerful unsupervised learning technique. The clustering process categorizes pixels into distinct groups based on their intensity values, effectively partitioning the image into K clusters.
To address potential artifacts and refine the segmentation output, a post-processing step is introduced. This involves the application of a binary mask generated from the clustered results. The mask is further refined by eliminating small objects or noise through morphological operations, enhancing the accuracy and coherence of the segmented regions.
1. Image-based segmentation serves as the foundational method for digitally converting leaf images into multiple distinct regions.
2. The partitioning is achieved using K-means clustering with three clusters.
3. The first cluster contains images representing normal results, while the second cluster signifies abnormal results.
4. Consideration is given to the diversity of category feature sets.
5. Centroids are calculated using Euclidean distance from the initial cluster, with data points employed in an iterative algorithm.
6. The distance is utilized to determine the cluster centroid across the defined clusters.
7. Let 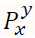 represent the clustered input pixel and denote the centre of the cluster.
represent the clustered input pixel and denote the centre of the cluster.
8. Initialize the number of clusters and their centres.
9. Calculate the Euclidean distance for each leaf-based pixel using the formula

10. Assign the nearest center for every pixel based on the calculated distance.
11. Calculate the new position using the expression
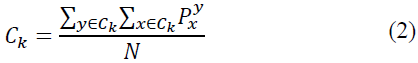
12. Repeat the process until the error value reaches a satisfactory level and Plot the relevance of the image.
This segmentation technique effectively isolates features relevant to mango leaf analysis, laying the foundation for subsequent stages such as feature extraction and disease classification.
3.3. Wavelet-Based Image Refinement
After segmentation, a wavelet-based refinement step is applied to enhance disease-related texture information in mango leaf images. In this study, the Daubechies (db4) wavelet was selected due to its proven effectiveness in capturing localized texture variations in agricultural imagery. A one-level discrete wavelet decomposition was performed on the segmented leaf image.
At each level of decomposition, the image is separated into four sub-bands: approximation (LL) and detail components (LH, HL, HH), corresponding to low-frequency and high-frequency information, respectively. The LL sub-band, which preserves the dominant structural characteristics of the leaf, was retained, while the high-frequency detail sub-bands (LH, HL, HH) were selectively emphasized to enhance disease-specific edge and texture patterns which is depicted in Figure 3.
The refined image was reconstructed using inverse wavelet transformation by combining the enhanced approximation and detail coefficients. This process suppresses background noise while amplifying discriminative texture variations caused by disease symptoms, resulting in an image that is better suited for feature extraction.
The refined image is subsequently used as input for HOG feature extraction, enabling more robust gradient-based representation of disease patterns prior to classification.
The post-processing stage significantly contributes to the fidelity of the image data, providing a clean and informative representation of mango leaves. This refined data, coupled with subsequent feature extraction using techniques like Histogram of Oriented Gradients (HOG) [21], forms the basis for robust machine learning-based classification, ultimately aiding in the accurate identification of healthy and Anthracnose-affected mango leaves.
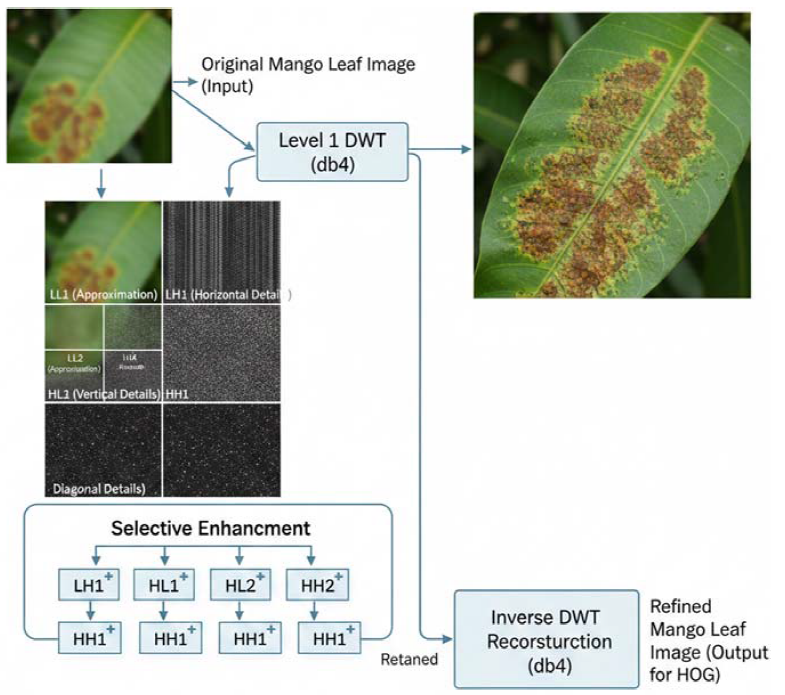
Figure 3. Wavelet based disease texture enhancement using wavelets.
3.4. Feature Extraction Using HOG
In the proposed research framework, a robust feature extraction process is integral to the comprehensive analysis of segmented mango leaf images. Feature extraction serves as a crucial intermediary step between segmentation and classification, aiming to distil relevant information that characterizes various aspects of the leaf structure. The feature extraction algorithm is designed to capture distinctive patterns and attributes within the segmented regions, facilitating the subsequent discrimination of healthy and various disease affected mango leaves.
The conventional Histogram of Oriented Gradients (HOG) feature extraction process consists of steps to capture image characteristics. Initially the input image is resized to a size of 64 × 64 pixels. Gamma normalization is then used to enhance the contrast of the image. Following that the magnitude and orientation of gradients are calculated for each pixel in the image. The image is divided into 8 × 8 pixel cells and a sliding window measuring 16 × 16 pixels moves, across these cells creating blocks that overlap four cells. Within each block trilinear interpolation is utilized to create a histogram based on orientations. These histograms are. Combined to create a one feature vector.
While effective traditional HOG feature extraction can be demanding in terms of computation. Result in a set of features. To tackle this issue researchers have suggested improvements such as using histogram techniques to Viola and Jones integral image approach. This method allows for histogram computation within any region of the image. Researchers like Yahia et al. Have utilized this method to speed up histogram calculation within each cell.
Due to its remarkable success in complex environments, the Histogram of Oriented Gradients (HOG) has garnered significant attention as an effective attribute extraction method and finds frequent utilization in the literature. Widely employed in object and pattern recognition, HOG exhibits high performance across diverse conditions [22].
The HOG method is known for extracting information regarding the distribution of target edges in local regions, providing a means to represent the target shape [22]. Employing HOG involves characterizing the orientation (θ) and magnitude (m) values of pixels in the image, making it of great interest to various fields. This method aims to define an image as a collection of local histograms.
Block normalization involves presenting the orientation histogram for each cell and subtracting larger blocks. The feature vector, denoted as v, represents the un-normalized histogram at point (i, j) of the block v (i, j). The feature vector undergoes normalization using the L2-norm method [22,23].
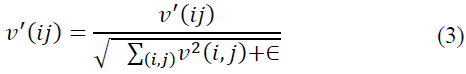
Here, ϵ is a very small normalization constant introduced to prevent division by zero [22].
Moreover there have been efforts to merge HOG features with extraction methods such, as Local Binary Patterns (LBP) Gabor filters and Discrete Wavelet Transform (DWT) [24]. The combination of these methods brings the benefit of acquiring traits, from each approach. Yet this blending could complicate the extraction process. Lead to feature dimensions requiring more time, for processing.
This methodological approach, leveraging HOG for attribute extraction and employing block normalization, contributes to the robustness and versatility of the technique, making it applicable across diverse domains. The resulting feature vector, encompassing information on gradient orientations and magnitudes, serves as a distinctive fingerprint characterizing the unique textural and shape attributes of mango leaves. This feature representation is subsequently utilized for training and evaluating machine learning models, contributing to the accurate classification of healthy and various types of affected mango leaves.
3.5. SVM-Based Classification
In our research, the classification stage serves as the core component of our automated system for mango leaf disease detection. This stage is instrumental in accurately categorizing mango leaf images into healthy and diseased classes, enabling timely intervention measures to mitigate disease spread and crop damage.
The extracted features are then fed into a Support Vector Machine (SVM) [25] classifier, a powerful machine learning algorithm known for its ability to handle high-dimensional feature spaces and nonlinear decision boundaries. SVM learns to discriminate between healthy and diseased mango leaves by identifying patterns and relationships within the feature space. Through the process of supervised learning, the SVM classifier optimizes its decision boundary to accurately separate the two classes.
Extensive experiments are conducted to evaluate the performance of the SVM classifier in classifying mango leaf diseases. The classifier is trained on a diverse dataset comprising images of mango leaves exhibiting various disease symptoms, including anthracnose, powdery mildew, and bacterial spot. Performance metrics such as accuracy, precision, recall, and F1 score are computed to assess the classifier's effectiveness in distinguishing between healthy and diseased leaves.
4.1. Data (Material)
Results are reported using the mango leaf image dataset consisting of four classes, namely healthy, powdery mildew, sooty mold, and anthracnose. A total of 2,000 images were used in the experiments, with 500 images per class.
The dataset was divided into training and testing subsets using a fixed 80:20 split, where 80% of the images were used for training and the remaining 20% were reserved exclusively for testing. Specifically, for each disease class, images were randomly partitioned such that no overlap occurred between the training and testing sets.
To prevent data leakage, dataset splitting was performed prior to any image preprocessing, including filtering, segmentation, wavelet transformation, and feature extraction. All preprocessing and feature extraction steps were applied independently to the training and testing subsets after the split.
Furthermore, 5-fold cross-validation was employed on the training set to optimize the SVM hyperparameters and evaluate model stability. The final performance metrics were reported on the held-out test set, which was never used during training or validation.
4.2. Evaluation Metrics
In the context of disease and healthy mango leaf classification for mango leaves, our proposed technique exhibits remarkable performance, achieving an outstanding accuracy [26] of 100.00%. Accuracy is computed as the ratio of correctly predicted instances (true positives and true negatives) to the total instances:
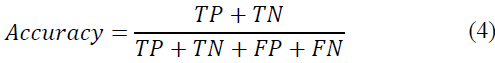
Where TP denotes true positives, TN denotes true negatives, FP denotes false positives, and FN denotes false negatives. Precision (P), recall (R), and F1 score (F1) serve as pivotal metrics for a comprehensive evaluation of model efficacy. These metrics are defined as follows:
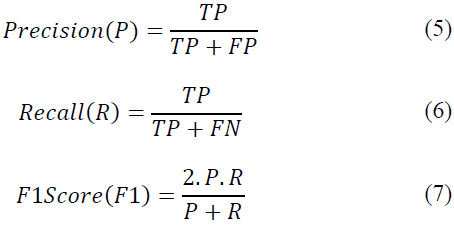
Here, TP signifies true positives, FP signifies false positives, and FN signifies false negatives. For Class 1 (Anthracnose-affected leaves), the precision, recall, and F1 score all achieve a perfect score of 1 (P=1, R=1, F1=1). These metrics affirm that every instance predicted as Anthracnose-positive by our model is indeed correct, with zero occurrences of false positives or false negatives. The exceptional precision, recall, and F1 score collectively underscore the reliability and accuracy of our proposed technique in Anthracnose detection.
4.3. Results of Experiments and Comparison
To assess the contribution of each component in the proposed framework, an ablation study was conducted by incrementally removing individual modules from the pipeline while keeping the remaining configuration unchanged (Table 1). The objective was to evaluate whether each processing stage contributes meaningfully to the final classification performance or if a simpler configuration could achieve comparable results.
Table 1. Accuracy comparison of different image processing and classification configurations.
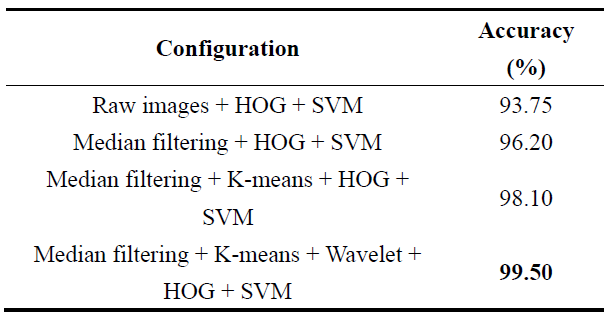

The results demonstrate that each module contributes incrementally to classification performance. Median filtering reduces noise and improves feature stability, while K-means segmentation isolates disease-relevant regions, leading to a noticeable accuracy improvement. Wavelet-based refinement further enhances multi-scale texture information, resulting in the highest classification accuracy when combined with HOG feature extraction and SVM classification. To evaluate the contribution of wavelet-based refinement, classification performance was compared with and without the wavelet decomposition step. Experimental results indicate that incorporating wavelet-based refinement improves classification accuracy and stability compared to using raw segmented images alone. These findings confirm that the complete pipeline is necessary to achieve optimal performance. All experiments were performed using the same training–testing split and SVM classifier to ensure fair comparison.
In evaluating the performance of our proposed technique, Conducted a comparative analysis with existing methods as presented in the literature. Table 2 summarizes the results of our technique alongside selected state-of-the-art methods.
4.4. Visualization of Results
To further illustrate the performance of the proposed framework, multiple visualization techniques are employed, including confusion matrices and accuracy bar plots.
The confusion matrix in Figure 4 provides a detailed class-wise evaluation by illustrating the relationship between actual and predicted labels for each disease category. Strong diagonal dominance in the matrix indicates high classification accuracy and minimal misclassification among visually similar disease classes.
Accuracy bar plots are used to visually compare the classification performance of the proposed method with existing approaches and ablation configurations in Figure 5. These plots offer an intuitive representation of performance improvements achieved through each stage of the proposed pipeline.
Table 2. Comparative analysis with existing techniques.
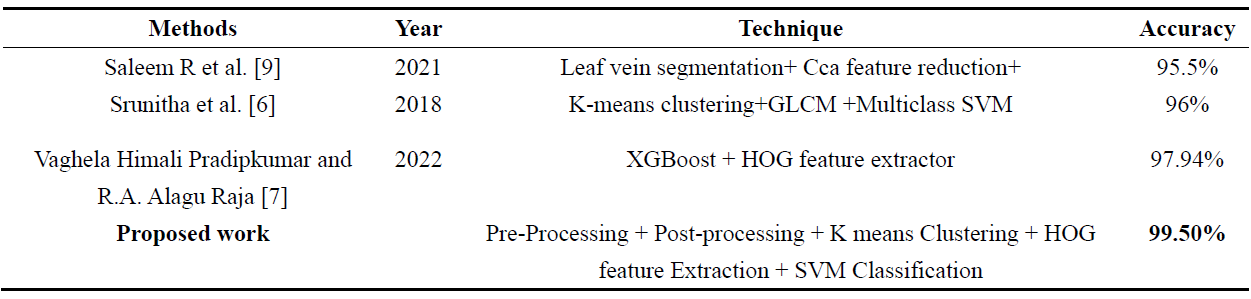

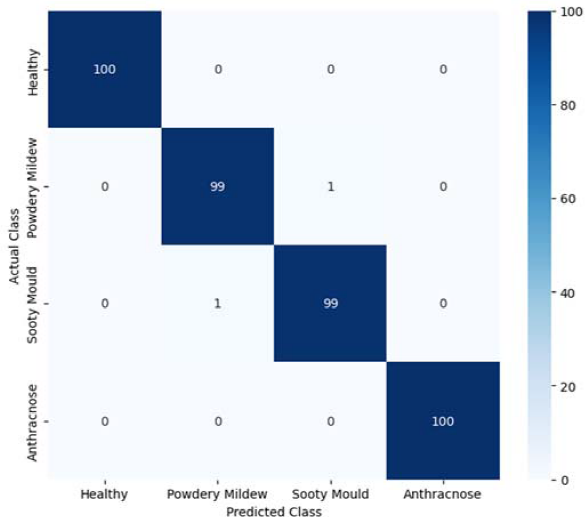
Figure 4. Confusion matrix illustrating the classification performance.
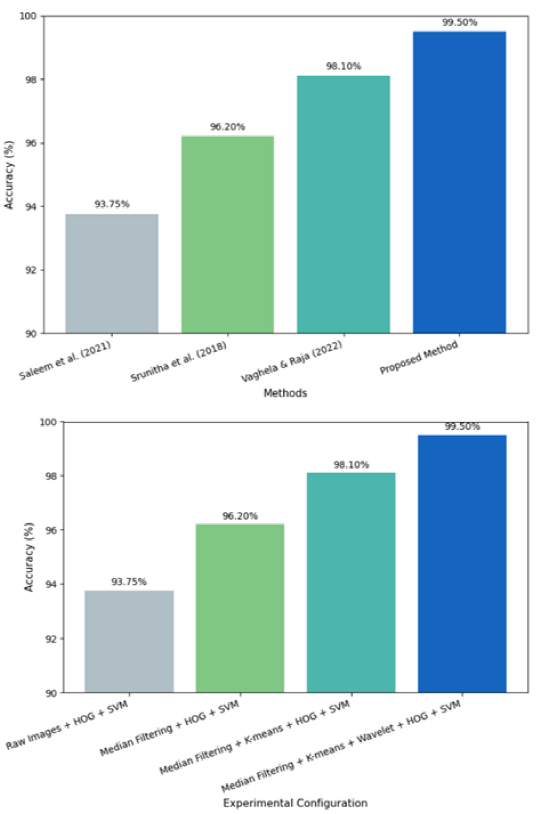
Figure 5. Comparative Accuracy Analysis of Mango Leaf Disease Classification Methods.
4.5. Discussion
The experimental results confirm the effectiveness of the proposed hybrid framework for mango leaf disease classification. The ablation study demonstrates that each module contributes incrementally to performance, with segmentation and wavelet-based feature enhancement playing a key role in improving disease-specific texture representation. The integration of HOG features with an SVM classifier enables robust class discrimination while maintaining computational efficiency.
Compared with existing methods, the proposed approach achieves superior accuracy without relying on deep learning architectures or large-scale datasets. This makes the framework more interpretable and suitable for practical agricultural applications. Overall, the results validate that the complete pipeline is necessary to achieve optimal performance and provides a reliable solution for automated mango leaf disease detection.
In summary, our proposed methodology showcases a robust framework for automated detection and classification of Anthracnose in mango leaves. Leveraging K-means segmentation, wavelet decomposition, and Histogram of Oriented Gradients (HOG) feature extraction, our approach achieves a remarkable 99.50% accuracy in distinguishing healthy from diseased leaves. The Support Vector Machine (SVM) with a polynomial kernel demonstrates high precision, recall, and F1 Score metrics, validating the efficacy of our method. This research not only contributes to automated plant disease detection but also holds potential for practical applications in precision agriculture, facilitating early intervention strategies for sustainable mango cultivation practices. Further exploration and refinement of this approach could lead to significant advancements in crop management and agricultural technology.
Author Contributions: Conceptualization, S.N. and M.G.R.; methodology, S.N.; software, S.N.; validation, S.N. and M.G.R.; formal analysis, S.N.; investigation, S.N.; resources, M.G.R.; data curation, S.N.; writing—original draft preparation, S.N.; writing—review and editing, S.N. and M.G.R.; visualization, S.N.; supervision, M.G.R.; project administration, M.G.R. All authors have read and agreed to the published version of the manuscript.
Conflicts of Interest: The authors declare no conflicts of interest.
Khairnar, K.; Goje, N.S. Image Processing Based Approach for Diseases Detection and Diagnosis on Cotton Plant Leaf. In Techno-Societal 2018; Pawar, P. M., et al., Eds.; Springer: Cham, Switzerland, 2020; pp. 57–65. DOI: 10.1007/978-3-030-16848-3_6.
Rao, U.S.; Swathi, R.; Sanjana, V.; Arpitha, L.; Chandrasekhar, K.; Chinmayi; et al. Deep Learning Precision Farming: Grapes and Mango Leaf Disease Detection by Transfer Learning. Global Transitions Proc. 2021, 2, 535–544.
Rehman, M.Z.U.; Ahmed, F.; Khan, M.A.; Tariq, U.; Jamal, S.S.; Ahmad, J.; et al. Classification of Citrus Plant Diseases Using Deep Transfer Learning. Comput. Mater. Contin. 2022, 70, 1401–1417.
Webster, C.; Ivanov, S. Robotics, Artificial Intelligence, and the Evolving Nature of Work. In Digital Transformation in Business and Society; Springer: New York, NY, USA, 2020; pp. 127–143.
Harakannanavar, S.S.; Rudagi, J.M.; Puranikmath, V.I.; Siddiqua, A.; Pramodhini, R. Plant Leaf Disease Detection Using Computer Vision and Machine Learning Algorithms. Global Transitions Proc. 2022, 3, 305–310.
Srunitha, K.; Bharathi, D. Mango Leaf Unhealthy Region Detection and Classification. In Computational Vision and Bio Inspired Computing; Hemanth, D., Smys, S., Eds.; Lecture Notes in Computational Vision and Biomechanics, Vol. 28; Springer: Cham, Switzerland, 2018; pp. 422–436. https://doi.org/10.1007/978-3-319-71767-8_35.
Pradipkumar, V.H.; Alagu Raja, R.A. Automatic Identification of Tree Species from UAV Images Using Machine Learning Approaches. J. Indian Soc. Remote Sens. 2022, 50, 2447–2464.
Mia, M.R.; Roy, S.; Das, S.K.; Rahman, M.A. Mango Leaf Disease Recognition Using Neural Network and Support Vector Machine. Iran J. Comput. Sci. 2020, 3, 241–250. DOI: 10.1007/s42044-020-00057-z.
Saleem, R.; Shah, J.H.; Sharif, M.; Yasmin, M.; Yong, H.-S.; Cha, J. Mango Leaf Disease Recognition and Classification Using Novel Segmentation and Vein Pattern Technique. Appl. Sci. 2021, 11, 11901. https://doi.org/10.3390/app112411901.
Salma, N.; Madhuri, G.R. Advancing Remote Sensing: A Unified Deep Learning Approach with Pretrained and Custom Architectures for High-Precision Classification. Phys. Scr. 2024, 99, 116012.
Rehman, M.Z.U.; Ahmed, F.; Khan, M.A.; Tariq, U.; Jamal, S.S.; Ahmad, J.; et al. Classification of Citrus Plant Diseases Using Deep Transfer Learning. Comput. Mater. Contin. 2022, 70, 1401–1417.
Singh, U.P.; Chouhan, S.S.; Jain, S.; Jain, S. Multilayer Convolution Neural Network for the Classification of Mango Leaves Infected by Anthracnose Disease. IEEE Access 2019, 7, 43721–43729.
Sujatha, R.; Chatterjee, J.M.; Jhanjhi, N.Z.; Brohi, S.N. Performance of Deep Learning vs Machine Learning in Plant Leaf Disease Detection. Microprocess. Microsyst. 2021, 80, 103615. DOI: 10.1016/j.micpro.2020.103615.
Usharani, M.; Shwetha, N.; Soundarya, Y. Detection of Pest in Agricultural Crops by SVM Algorithm. Int. J. Eng. Res. Technol. 2019, 8, 1–4.
Pradipkumar, V.H.; Alagu Raja, R.A. Automatic Identification of Tree Species from UAV Images Using Machine Learning Approaches. J. Indian Soc. Remote Sens. 2022, 50, 2447–2464.
Khan, S.B.J.; Peng, Z.; Kamal, M.M.; Mohamed, H.G.; Kharma, Q.M.; Sheraz, M.; et al. STRACK: Robust Tracking of Small Objects in Low-Light Conditions. IEEE Access 2025, 13, 12345–12360.
Baz, J.K.S.; Zhang, P.; Kamal, M.M.; Mohamed, H.G.; Sheraz, M.; Chuah, T.C. HAMOT: A Hierarchical Adaptive Framework for Robust Multi-Object Tracking in Complex Environments. Comput. Model. Eng. Sci. 2025, 145, 123–145.
Kamal, M.M.; Abideen, S.Z.U.; Al-Khasawneh, M.A.; Momani, A.M.; Mostafa, H.; Atoum, M.S.; et al. Meyer Wavelet Transform and Jaccard Deep Q Net for Small Object Classification Using Multi-Modal Images. Comput. Model. Eng. Sci. 2025, 144, 3053–3083.
Salma, N.; Madhuri, G.R. Synergistic Use of Convolutional Neural Networks and Support Vector Machines for Mango Leaf Disease Diagnosis. Sci. J. King Faisal Univ. Basic Appl. Sci. 2025.
Camargo, A.; Smith, J.S. Image Pattern Classification for the Identification of Disease Causing Agents in Plants. Comput. Electron. Agric. 2009, 66, 121–125.
Kiran, S.M.; Chandrappa, D.N. Plant Leaf Disease Detection Using Efficient Image Processing and Machine Learning Algorithms. J. Robot. Control 2023, 4, 846–854. DOI: 10.18196/jrc.v4i6.20342.
Adeel, A.; Khan, M.A.; Sharif, M.; Azam, F.; Shah, J.H.; Umer, T.; et al. Diagnosis and Recognition of Grape Leaf Diseases: An Automated System Based on a Novel Saliency Approach and Canonical Correlation Analysis Based Multiple Features Fusion. Sustain. Comput. Inform. Syst. 2019, 24, 100349.
Anantrasirichai, N.; Hannuna, S.; Canagarajah, N. Towards Automated Mobile-Phone-Based Plant Pathology Management. arXiv 2019. arXiv:1912.09239.
Rakshitha, G.; Yashaswini, B.M.; Sneha, M.; Singh, A.; Sonu, C. Plant Disease Detection Using Machine Learning. Int. J. Adv. Res. Ideas Innov. Technol. 2020, 6, 427–429.
Jaisakthi, S.M.; Mirunalini, P.; Thenmozhi, D.; Vatsala. Grape Leaf Disease Identification Using Machine Learning Techniques. In 2019 International Conference on Computational Intelligence in Data Science(ICCIDS), Chennai, India, 21–23 Feb. 2019; pp. 1–6.
Balasundaram, A.; Pradeep, K.V.; Sandhya, S. An Extensive Study on Disease Prediction in Mango Trees Using Computer Vision. Ann. Rom. Soc. Cell Biol. 2021, 25, 1895–1905.